Informed and Effective Treatments for Your Patient
What is your greatest challenge when a patient walks through your door?
If the patient is a newly diagnosed patient, for most oncologists, it is not being certain of response to standard care therapy. If it is a patient for whom standard care has already failed, it is determining what treatment options may be available for the patient. If the patient has been diagnosed with an indication that is poorly responsive to treatment, it is deciding the next course of action. You are dealing with situations that limit your options because of challenges in predicting responsiveness to therapy, when time is limited.
The challenge of matching therapies to individual patients is complex. What has traditionally been diagnosed as a single indication via biomarkers, histology and cell morphology is now understood to be multiple disease subtypes in the context of biochemical pathways. As a consequence, the ‘one mutation, one drug’ paradigm, has proven to be of limited value for treatment decisions. The emergence of NGS helped identify broad tumor genomic profiles that provide additional data. However, the clinical value of NGS reports has been disappointing. In addition to an already confounding challenge, the physician is further burdened with indiscernible information.
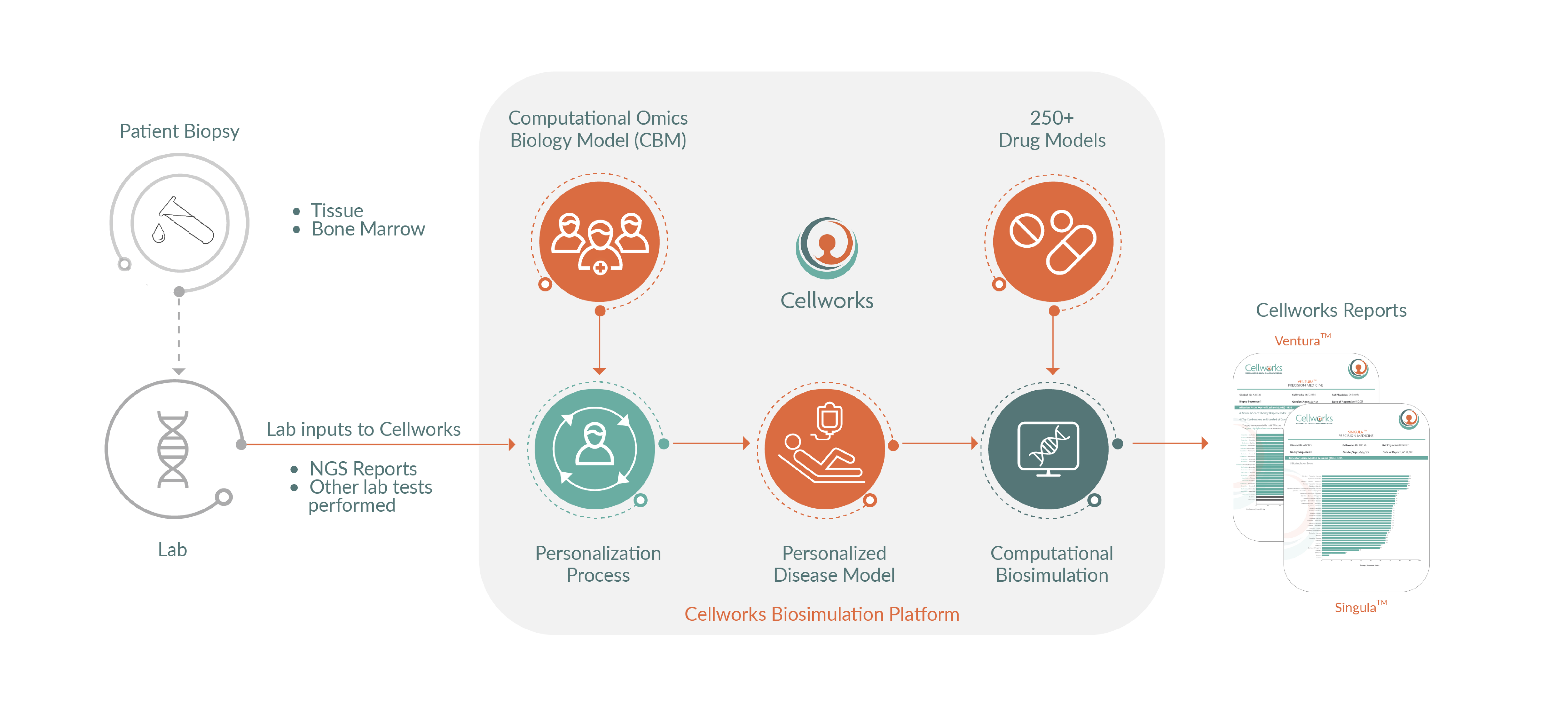
Cellworks assists interpretation of heterogeneity in a patient’s tumor genomic profiling using computational mathematics and biosimulation.The patient disease model is created to understand the biochemical molecular pathways and clinical outcomes of treatment on disease phenotypes. Cellworks Therapy Response Index (TRI) reports provide you with actionable suggestions, each with a transparent causative rationale.
SINGULATM Standard Care Response:
Cellworks SingulaTM predicts personalized response to Standard Care therapies for front-line patients.
Download Sample Report for SingulaTM
By predictively separating responders from non-responders, the physician has actionable decision making ability for optimized therapy. This can save vital time for the patient as well as control treatment costs. Say you, the physician, are presented with an MDS patient who has TET1, TET2, MTHFR, MTRR, TP53, APC, AXIN2 and ASXL1 mutations and a complex cytogenetic profile with loss of various chromosomal segments. SingulaTM can help guide you whether the patient with this set of genomic aberrations is likely to respond to an MDS standard care drug like Azacitidine or Lenalidomide.
Show More
Show Less
VENTURATM Ideal Combination:
Cellworks VenturaTM predicts and ranks personalized response to combinations of FDA-approved drugs including off-label and non-oncology drugs for refractory patients.
Download Sample Report for VenturaTM
After the standard care therapies are exhausted, or after the patient has relapsed, the current protocol is to send the patient to clinical trials or to palliative care. Cellworks offers another option. VenturaTM informs you, the physician, of the likely ideal combination of all FDA-approved drugs to produce the response in the patient. In determining this optimal combination, on-label oncology drugs, off-label oncology drugs and non-oncology drugs are all investigated before a recommendation is made.
Say you, the physician, are presented with an AML patient who has HER2, FGFR4, IL4R, IL6R, IL7, TET1, DNMT3L mutations, and various other genes have copy number variations and are amplified (ACLY, NRG1, FGFR1, FGFR3, LYN) or deleted (TET3). VenturaTM will help guide you as to which combination of FDA-approved drugs this AML patient is likely to respond to.
Show More
Show Less
Validated. Precisely.
Cellworks’ biosimulation predictions have been validated through cell-line studies, animal studies, human ex vivo experiments, retrospective clinical studies and prospective clinical trials, with biomarker and phenotype trends in lab and clinical settings.
Cellworks has:
- Participated in more than 50 global scientific collaborations
- Conducted over 3000 prediction validations
- Produced over 100 peer-reviewed papers and abstracts
Engage In Improving Patient Outcomes
Start an engagement with Cellworks.
Participate in this paradigm shift. There are ongoing PI-initiated clinical trials that compare predictions with outcomes of cancer treatments for Multiple Myeloma (MM), Myelodysplastic Syndrome (MDS), Acute Myeloid Leukemia(AML), Myelofibrosis(MF), Glioblastoma(GBM), Early T-Cell Precursor Acute Lymphoblastic Leukemia(ETP-ALL), Waldenstrom Macroglobulinemia(WM), and Radiation Oncology.