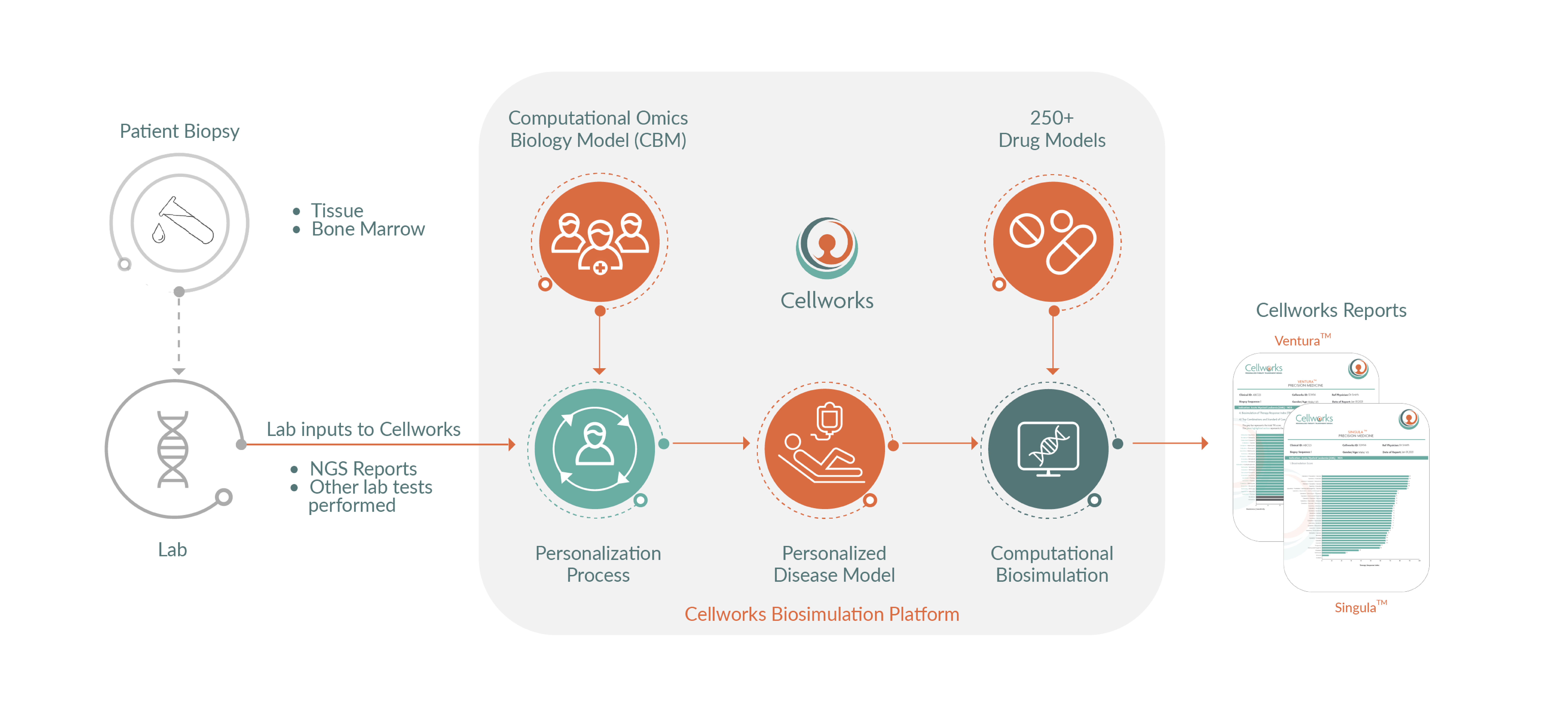
The Cellworks Biosimulation Platform
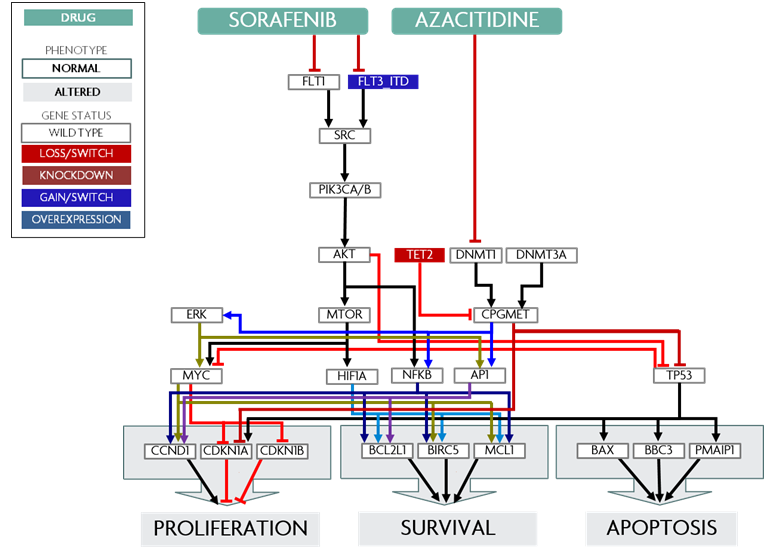
The Cellworks Platform biosimulates how a patient’s unique cancer will respond to therapies prior to treatment and identifies novel drug combinations for treatment-refractory patients. Therapy biosimulation is the ability to predict the phenotype response of a human cell to an external stimulus, such as a drug ligand or radiation. We utilize this technology, to determine a patient’s Therapy Response Index, or TRI. A treatment-naïve patient’s TRI determines whether the patient will respond to standard care treatment. For relapsed or refractory patients, TRI determines the ideal combination of FDA approved drugs to which the patient will respond.
Computational Omics Biology Model (CBM)
The biosimulation process begins with the Cellworks Computational Omics Biology Model (CBM), a groundbreaking in silico model of a healthy human cell comprised of a network of 4,000+ human genes, 30,000+ molecular species and 100+ signaling pathways. Using a patient’s cytogenetic and molecular data as input to the CBM, the Cellworks Platform generates a personalized disease model for each patient. The platform then dynamically simulates the phenotype response of the patient’s disease model to various therapies.
250+ Drug Models
There are more than 250 drug models within the Cellworks platform, including a quantitative model of drug targets, binding affinities, reaction rate constants coupled with pharmacokinetic and drug economic data.
AI-Driven Analytics
Machine learning techniques are used within the platform to understand the behavior of biochemical pathways and represent them mathematically in the model. Smaller pathways are combined into larger macro interactions using a manual curation approach.
Scalable, Cloud-based Infrastructure
The cloud-based infrastructure can run millions of prediction biosimulations in parallel and has a demonstrated ability to handle thousands of patients per day. The platform can scale differential equations and other mathematical structures at the rate of millions per second per biosimulation.
Transparent Design
Unlike conventional ex vivo testing and machine learning technologies that operate in a ‘black box’ mode with little explanatory power, Cellworks describes with full causality, the mechanism by which a therapy works for a given patient’s disease.