ASCO 2020 : myCare Clinical Studies Show Cellworks Singula™Achieves Superior Therapy Response Predictions for Brain Cancer Patients
Cellworks Multi-omics Biosimulation Delivers GBM and LGG Survival Predictors Above and Beyond Physician Prescribed Treatments
SOUTH SAN FRANCISCO, Calif., May 29, 2020 -- Cellworks Group, Inc., a world leader in Personalized Medicine in the key therapeutic areas of Oncology and Immunology, today announced results from the
myCare-009-03 and myCare-009-04 clinical studies,
which demonstrate that Cellworks Singula™ is a superior predictor of progression-free survival (PFS) and overall survival (OS) in Glioblastoma (GBM) and Low-Grade
Glioma (LGG) patients, compared to physician prescribed treatments.
The results from myCare-009-03 and myCare-009-04 clinical studies will be featured as poster presentations with comments from Dr. Manmeet Ahluwalia, MD, Cleveland Clinic, Brain Tumor and Neuro-Oncology Center as part of the ASCO20 Virtual Scientific Program May 29 – May 31, 2020 and available as online abstracts 2519 and 2569, respectively. ASCO20 registrants can also listen to a discussant commentary in the Central Nervous System Tumors Poster Session Track titled, Identifying and Validating Novel Biomarkers in Glioma, with Dr. Karla Ballman, Chief of Biostatistics and Epidemiology, Weill Cornell Medical College, in which the myCare-009-03 clinical study will be discussed.
“Biosimulating therapy responses using patient-specific multi-omics data is an important advancement for GBM and LGG patients,” said Dr. Patrick Wen, MD, Director, Center For Neuro-Oncology, Dana-Farber Cancer Institute; Professor, Neurology, Harvard Medical School; and co-Primary Investigator for myCare-009-03 and myCare-009-04 clinical studies. “These studies underscore the benefits of moving beyond a sole reliance on NGS reports for prescribing cancer treatments. The predictions in these studies delivered precise therapy response information at an individual patient level – fulfilling on the promise of Personalized Medicine.”
Outcomes for GBM and subsets of LGG patients have remained poor despite using cytogenetic and molecular-risk stratification along with Precision Medicine. Therapy selections for GBM and LGG patients are typically based on next-generation sequencing (NGS) reports and information that only considers a single aberration while ignoring many molecular pathways as well as other patient-specific multi-omics data, which can be used to inform a more effective therapy selection.
“In order to improve outcomes for GBM and LGG patients, we need to understand the effects of multiple aberrations at an individual patient level, which requires analysis beyond that of current NGS reports,” said Dr. Ahluwalia, MD, Cleveland Clinic, Brain Tumor and Neuro-Oncology Center and Primary Investigator for myCare-009-03 and myCare-009-04 clinical studies. “These Cellworks myCare studies illustrate that by using multi-omic data and biosimulating GBM and LGG patient therapy responses with the Cellworks Biosimulation Model (CBM), we gain predictive information above and beyond physician prescribed treatments that currently rely only on NGS reports.”
Cellworks Singula™ therapy response predictions are generated through extensive biosimulation of a personalized patient disease model based on the patient’s multi-omics data. Utilizing an in-silico model of thousands of genes, Cellworks Singula™ analyzes the downstream pathway impact of genomic, proteomic, transcriptomic and epigenomic aberration information on a patient’s disease. These downstream effects generate phenotypic impact, which are calculated against specific drugs or drug combinations to determine treatment efficacy. The Cellworks biosimulation platform can predict an individual patient’s therapy response prior to receiving the treatment, thereby reducing the patient risks and payer costs of unsuccessful treatments and ultimately saving lives.
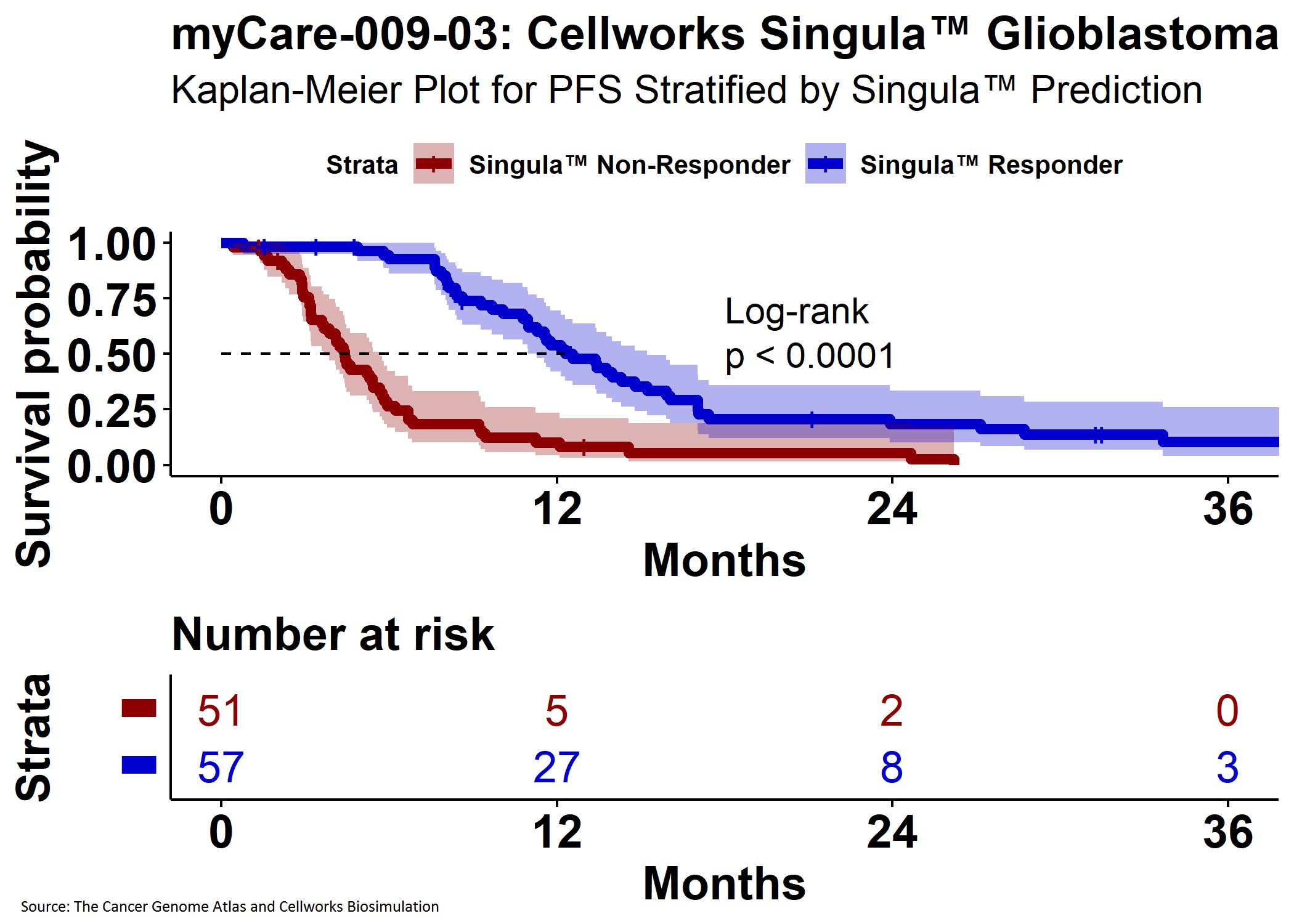
myCare-009-03 Clinical Study
In this prospective retrospective study, an independent randomly selected cohort of 109 GBM patients aged 17 to 83 years and treated with physician prescribed therapies were selected. Patient multi-omics data from The Cancer Genome Atlas (TCGA) was used along with information from PubMed to generate protein interaction network activated and inactivated disease pathways. The physician prescribed therapy for each patient was biosimulated to calculate the quantitative drug effect, using a composite GBM disease inhibition score based on specific phenotypes, while blinded to clinical response.
Study results show Cellworks Singula™ delivered predictions with 91.2% positive predictive value (Exact 95% CI: 80.7%, 97.1%) and 90.4% negative predictive value (95% CI: 79.0%, 96.8%) for clinical response as defined as either a complete response or partial response. In the study, Cellworks Singula™ performed with 91.2% sensitivity (95% CI: 80.7%, 97.1%) and 90.4% specificity (95% CI: 79.0%, 96.8%). The results also indicate that Cellworks Singula™ is a superior predictor of OS (HR = 1.852, p = 0.0070) and PFS (HR = 4.0333, p = < 0.0001) compared to physician prescribed treatments using NGS reports alone.
myCare-009-04 Clinical Study
An independent cohort of 137 LGG patients aged 14 to 73 years and treated with physician prescribed treatments were randomly selected for this prospective retrospective study. Patient multi-omics data from The Cancer Genome Atlas (TCGA) was used along with information from PubMed to generate protein interaction network activated and inactivated disease pathways. The physician prescribed treatment was biosimulated for each patient to calculate the quantitative drug effect, using a composite LGG disease inhibition score based on specific phenotypes, while blinded to clinical response.
Study results show CellworksSingula™ delivered 89.3% positive predictive value (Exact 95% CI: 81.7%, 94.6%) and 82.4% negative predictive value (95% CI: 65.5%, 92.3%) for clinical response as defined as either a complete or partial response. In the study, CellworksSingula™ performed with 93.9% sensitivity (95% CI: 87.1%, 97.7%) and 71.8% specificity (55.1%, 85.0%). Results from the study demonstrate that Cellworks Singula™ is a high predictor of OS (HR = 2.970, p = 0.0013), a significant independent predictor of PFS (HR = 3.707, p = < 0.0001) in LGG patients and provides predictive information above and beyond physician prescribed treatment. Cellworks Singula™ also correctly identified non-responders to physician prescribed treatments and provided alternative therapy selections for all 34 disease progressors.
“These studies demonstrate significant progress in the journey to increase survival rates for brain cancer patients,” said Yatin Mundkur, CEO of Cellworks. “Cellworks Singula™ enables oncologists to deeply understand which therapies will be efficacious for individual GBM and LGG patients and to avoid prescribing ineffective therapies that waste precious time.”
About Cellworks Group
Cellworks Group, Inc. is a world leader in Personalized Medicine in the key therapeutic areas of Oncology and Immunology. Using innovative multi-omics modeling, computational biosimulation and Artificial Intelligence heuristics, Cellworks predicts the most efficacious therapies for patients. The Cellworks unique biosimulation platform is a unified representation of biological knowledge curated from heterogeneous datasets and applied to finding cures. Backed by Sequoia Capital and Artiman Ventures, Cellworks has the world’s strongest trans-disciplinary team of molecular biologists, cellular pathway modelers and software technologists working toward a common goal – attacking serious diseases to improve the lives of patients. The company is based in South San Francisco, California and has a research and development facility in Bangalore, India. For more information, visit www.cellworks.life and follow us on Twitter @cellworkslife.
All trademarks and registered trademarks in this document are the properties of their respective owners.
Media Contacts:
Barbara Reichert
Reichert Communications, LLC
Barbara@reichertcom.com
415-225-2991
Michele Macpherson, Chief Business Officer
Cellworks Group, Inc.
Michele.macpherson@cellworksgroup.com




